Rhododendron colemanii: A New Species of Deciduous Azalea from the Coastal Plain of Alabama and Georgia
Wenyu Zhou, Taylor Gibbons, Loretta Goetsch and Benjamin Hall
Department of Biology, University of Washington, Seattle, Washington
Thomas Ranney
Department of Horticultural Science
Mountain Horticultural Crops Research and Extension Center
North Carolina State University, Fletcher, North Carolina
Ron Miller
Pensacola, Florida
abstract . A new species of deciduous azalea, Rhododendron colemanii R. Miller (Red Hills azalea), from the upper Coastal Plain of Alabama and Western Georgia, USA, has been characterized based on morphology, flowering phenology, habitat, native range, position in a molecular phylogeny, and genome size. Though specimens have previously been confused with Rhododendron alabamense Rehder or its hybrids, phylogenetic relationships based on sequences of the nuclear gene RPB2-I gene showed R. colemanii to be a coherent species in a clade of section Pentanthera G. Don along with R. luteum Sweet and the Southeastern U.S. azaleas R. atlanticum (Ashe) Rehder, R. austrinum (Small) Rehder, and R. calendulaceum (Michaux) Torrey. Flow cytometry showed R. colemanii to be, like the other azaleas in this clade, tetraploid. A detailed morphological description and Latin diagnosis are provided.
For the rhododendron community, as for the non-horticultural public, the name "azalea" is, unfortunately, a term shrouded in ambiguity. It has been applied to two sizable groups of
Rhododendron
species that bear little resemblance to one another. There are approximately 100 species of azaleas included in the evergreen sections
Tsutsusi
and
Brachycalyx
(Chamberlain and Rae, 1990), while in the deciduous section
Pentanthera
there are 16 to 18 species also commonly known as azaleas. Besides these well-delineated groups, a few additional species are also called azaleas, as exemplified by
R. vaseyi
Gray,
R. nipponicum
Matsumura,
R. schlippenbachii
Maximowicz,
R. albrechtii
Maximowicz, and
R. albiflorum
Hooker. These five deciduous azalea species share certain morphological traits with
Tsutsusi
azaleas, and recent DNA studies provided evidence that they are related to
Tsutsusi
azaleas (Goetsch et al., 2005).
Pentanthera
azaleas, on the other hand, were found by DNA analysis to be the sister group to
Hymenanthes
rhododendrons (Kurashige et al. 2001, Goetsch et al., 2005).
In this article, we present findings from two types of DNA analyses of
Pentanthera
azaleas: relative genome size and gene sequence determinations. These allow us to infer, respectively, the ploidy number of each species and an evolutionary tree that relates the different species to one another. This research began as a collaborative project to find out whether a deciduous azalea from the Red Hills region of Alabama and Georgia, initially collected and propagated by S. D. Coleman, Sr., is a new species. That turned out to be the case. (For a detailed history of the Red Hills azalea, see Miller and Yeatts, 2008.) As the scope of the investigation widened to include all species in section
Pentanthera
, Clarence Towe put the University of Washington researchers into communication with Tom Ranney of North Carolina State University. In discussions with Ben Hall, Clarence had noticed a surprising correlation. A group of seven species, including the Red Hills azalea, had DNA sequences that were similar to one another, but markedly different from the other eleven
Pentanthera
azalea species. This group of seven included all five of the
Pentanthera
species that Ranney's lab had found to be tetraploid, while all eleven in the other group are diploid (Jones et al. 2007). Further investigation of this correlation has led us to conclude that the tetraploid
Pentanthera
species have long been evolving as a separate lineage, reproductively separated from all but two of the known diploid species.
Materials and Methods
Taxon Sampling
. The species and clones studied for molecular and cytogenic components included all previously described
Pentanthera
azaleas (Kron, 1993, Kron and Creel, 1999), the Red Hills azalea, and the related species
R. canadense
(L.) Torrey. The molecular analyses also included three, more distantly related species, which we used as outgroups to root our evolutionary trees:
R. vaseyi
,
R. minus
Michaux and
R. maximum
L. Flower buds were collected from individual plants for DNA sequencing and relative genome size determination. Except for the Red Hills samples, which were collected either from the wild or from Callaway Gardens, Georgia, USA, the remaining taxa were collected from the Rhododendron Species Foundation Botanical Garden, Federal Way, Washington, USA. The plants, grown there from wild-collected seed, had morphological characteristics consistent with their published descriptions (Kron, 1993). Morphological measurements of the Red Hills azalea were completed on representative samples from throughout its geographical range.
Flow Cytomet ry and Ploidy Determination
.
Holoploid, 2C DNA contents (i.e., DNA content of the entire non-replicated, chromosome complement irrespective of ploidy level) were determined via flow cytometry following the methods of Jones et al. (2007). Genome sizes were determined by comparing mean relative fluorescence of each sample with an internal standard,
Pisum sativum
L. 'Ctirad', with a genome size of 8.76 ρg (Greihuber et al., 2007) and calculated as: 2C DNA content of sample = 8.76 ρg x mean fluorescence value of sample/mean fluorescence value of standard). Ploidy levels from these samples were determined based on the established relationship between relative DNA content and ploidy level in Rhododendron as reported by Jones et al. (2007).
DNA Extraction, Sequencing and Phylogenetic Analysis
.
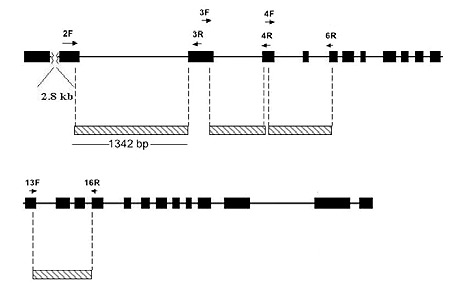
The methods used for extracting DNA from leaf or flower bud tissue, DNA amplification by the Polymerase Chain Reaction (PCR) procedure, DNA sequencing and phylogenetic analysis are all described in Goetsch et al. (2005). The gene sequences studied are those in the RPB2-I gene (Figure 1), which codes for a subunit of the enzyme RNA Polymerase II.
|
|
Figure 1. Structure of the Rhododendron RPB2-I Gene. The gene has
25 exons (black blocks) that make up the protein coding region. Between these are introns, indicated by a black line. DNA sequences used in this study are marked by cross-hatched blocks. Except for exons 5, 14 and 15, most of the sequence analyzed comes from introns. The small arrows mark the positions of DNA primers used to amplify DNA for sequencing. |
Results
Flow Cytometry and Ploidy Determination
.
Relative genome sizes of the
Pentanthera
azaleas (Table 1) fell into one of two classes consistent with either diploids (ranging from 1.17 to 1.57 ρg, mean = 1.48) or tetraploids (ranging from 2.61 to 3.22 ρg, mean = 3.04), as established for
Rhododendron
by Jones et al. (2007). Although we used similar methods to Jones et al. (2007), we employed an updated estimate of genome size for the internal standard (
Pisum sativum
L. 'Ctirad') of 8.76 ρg (Greihuber et al., 2007), that uniformly resulted in slightly lower values.
In a previous study that included
Pentanthera
azaleas, Jones et al. (2007) confirmed the tetraploidy of
R. calendulaceaum
and
R. luteum
and further determined that
R. austrinum
and
R. atlanticum
were tetraploid species. All the other
Pentanthera
species they studied were predominantly diploid. (These correspond to Table 1, lines 3 to 13 in this paper.) The research findings we report here (Table 1) corroborate those of Jones et al. (2007) for the species mentioned above. In addition, we found
R. molle
(Blume) G. Don and
R. canadense
to be diploid (Table 1). In view of previous reports of tetraploid
R. canadense
(Ammal et al., 1950; Sax, 1930), it remains to be seen whether the diploid state is limited to the
R. canadense
clone we studied.
Determination of the ploidy for three different accessions of the Red Hills azalea showed that they were all tetraploids (Table 1). This raises the very interesting question of the evolutionary relationship of the Red Hills azalea to other tetraploid
Pentanthera
azalea species (Jones et al., 2007).
| Table 1. Relative genome sizes and ploidy levels of Pentanthera azaleas determined by flow cytometry. | |||
| Taxa | Accession - Source 1 | Relative 2C genome size 2 | Ploidy Level (x) |
| R. canadense | Hootman - RSF | 1.17 ± 0.00 | 2 |
| R. molle | 80/091 - RSF | 1.44 ± 0.02 | 2 |
| R. occidentale | 77/388 - RSF | 1.47 ± 0.01 | 2 |
| R. eastmanii | 2004/037 - RSF | 1.48 ± 0.07 | 2 |
| R. prinophyllum | 98/155 - RSF | 1.50 ± 0.03 | 2 |
| R. cumberlandense | 2004/059 - RSF | 1.50 ± 0.03 | 2 |
| R. prunifolium | 2003/322 - RSF | 1.50 ± 0.03 | 2 |
| R. periclymenoides | 76/292 - RSF | 1.51 ± 0.03 | 2 |
| R. viscosum | 2004/193 - RSF | 1.51 ± 0.01 | 2 |
| R. flammeum | 76/286 - RSF | 1.53 ± 0.02 | 2 |
| R. arborescens | 80/012 - RSF | 1.56 ± 0.01 | 2 |
| R. canescens | 76/278 - RSF | 1.56 ± 0.00 | 2 |
| R. alabamense | 81/119 - RSF | 1.57 ± 0.00 | 2 |
| R. luteum | 88/065 - RSF | 2.61 ± 0.03 | 4 |
| R. atlanticum | 73/010 - RSF | 3.02 ± 0.03 | 4 |
| R. calendulaceum | 76/292 - RSF | 3.09 ± 0.02 | 4 |
| R. colemanii | A (Coleman/Callaway-Towe) | 3.10 ± 0.06 | 4 |
| R. austrinum | 2006/041 - RSF | 3.11 ± 0.04 | 4 |
| R. colemanii | B (Grove Hill)-Towe | 3.16 ± 0.06 | 4 |
| R. colemanii | C (Fountain)-Towe | 3.22 ± 0.03 | 4 |
|
1
Sources: RSF - Rhododendron Species Foundation, Federal Way, Wash.; Clarence Towe, Walhalla, SC.
2 Values are mean 2C relative genome size ± SEM for two samples. |
|||
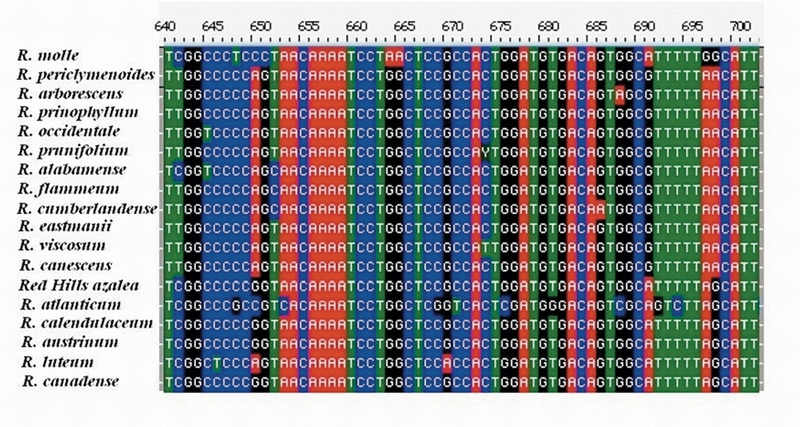
DNA Sequencing and Phylogenetic Analysis . The sequence data set for 18 Pentanthera species and three outgroups consisted of 3200 base pairs of aligned RPB2-I sequence for each, the vast majority of it from intron (non-coding) sequences (Figure 1). A short segment of the alignment is shown in Figure 2, where a number of base changes can be seen to subdivide the species into groups that differ in the base present at several positions. This set of aligned RPB2-I sequences was used to infer an evolutionary tree by two different approaches. The first, called maximum parsimony, uses the DNA data to compute the evolutionary tree with the smallest number of steps (mutational changes). The second method, maximum likelihood, minimizes the total combined length of all tree branches, including terminal ones. With the RPB2-I sequence data, both analytical methods produced the same tree (Figure 3). The outgroup species provide information about the root of the Pentanthera evolutionary tree. In this instance, the two clades of Pentanthera azaleas labeled A and B relate similarly to the outgroups. Thus, their DNA sequences do not indicate that either of them is clearly ancestral.
|
|
Figure 2. Aligned Sequences for a 62 base pair region within intron 2 of RPB2-I. Note the base differences between
groups of species at positions 641, 650, 691, and 698. Positions such as these serve to distinguish between clades in the phylogenetic analysis. A clade is a group of species that includes all descendants of one common ancestor. Clade and Monophyletic group are synonymous. |
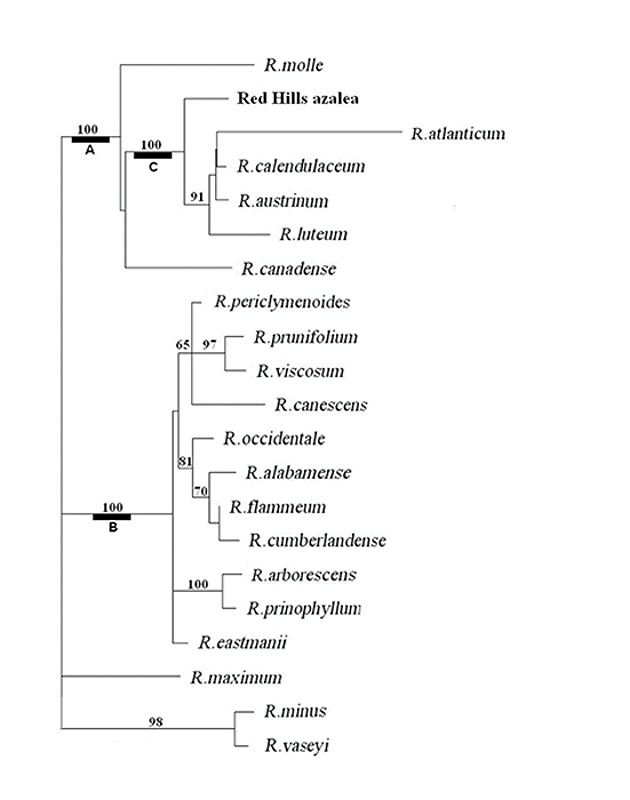
Each of the phylogenetic analyses provides, in addition to a tree that best fits the DNA data, useful information regarding the degree of certainty of various features of the tree. For the parsimony analysis, this measure of statistical support is called bootstrap support, shown as numbers adjacent to the nodes where two branches of the tree diverge (Figure 3). A bootstrap figure of 100 indicates the highest degree of certainty for that node, while 50 or less indicates low certainty and 70 or higher a reasonable degree of certainty. The most striking result of the parsimony analysis is that there is 100% support for the tree nodes that define clades A, B and C (Figure 3). This means that the evidence for these as distinct groups of genes is unequivocal. All the more remarkable is the fact that at the organismal level, the species corresponding to DNA clades A and C are homogeneous for ploidy, with all species in A being diploid and all in C being tetraploid. Rhododendron canadense and R. molle , the two species remaining outside of clades B and C, are more closely related to clade C than to clade B, suggesting a possible shared ancestry with the tetraploid azaleas. The phylogenetic position of the Red Hills azalea is within the C clade. Thus, not only is the Red Hills azalea tetraploid (Table 1), all of its close relatives are tetraploid, as well (Table 1). This clearly distinguished Red Hills from R. alabamense , which is diploid and is most closely related to other diploid species (Figure 3, Table 1).
|
|
Figure 3. Phylogenetic tree of
Pentanthera
azaleas. The tree shown is that from maximum
likelihood analysis. Branch lengths are proportional to the number of base changes on that branch. Maximum parsimony gave the same tree. Numbers indicate bootstrap support in the parsimony analysis. Authors of species not previously mentioned in this article: R. cumberlandense E. L. Braun, R. flammeum (Michaux) Sargent, R. periclymenoides (Michaux) Shinners, R. prinophyllum (Small) Millais, R. viscosum (L.) Torrey |
Discussion
The Red Hills Azalea
. This study analyzed DNA sequences of the RPB2-I gene to categorize the
Pentanthera
section of deciduous azaleas and demonstrated that the Red Hills azalea is closely allied to, but distinct from, the other
Pentanthera
species. Because the Red Hills azalea possesses the morphological characteristics of section
Pentanthera
, it must then be a new azalea species within this section.
Relationships within Section Pentanthera . At an early point in the evolutionary divergence of Pentanthera azalea species, two lineages separated - one containing the common ancestor of R. molle , R. canadense and clade C, the other being the common ancestor of R. occidentale (Torrey & Gray) Gray and all of the remaining Pentanthera species of the Eastern US. The lineages in question are RPB2-I gene lineages, which may or may not reflect relationships that apply across the entire genome. To explore these possibilities, further experiments, now in progress, will determine whether analyses of other regions of the Rhododendron genome recover the same evolutionary tree.
The Origin of Tetraploids, Ancient or Recent? The observation that hybrid Pentanthera azaleas are frequently seen in natural populations (Towe, 2004; Skinner, 1955) has led many to consider the possibility that tetraploid species may have originated recently in conjunction with interspecific hybridization, i.e., allopolyploidy (Li, 1957). One conceptually simple mechanism would be simply to have the pollen of one diploid species fertilize the ovum of another diploid species. Chromosomal doubling, either pre-or post-fertilization, could have led to stable, fertile (4x) hybrids having the diploid complement of chromosomes from each parent. The net result would be a new tetraploid species formed from diploid ancestors. The results we have obtained do not support this mechanism for the recent genesis of tetraploid Pentanthera species. None of the tetraploid species in clade C, including R. colemanii , contains an RPB2-I gene like that of any diploid species. The fact that these five species make up a monophyletic group, sharing a common ancestor unique only to them, has important implications. This group contains both R. luteum , native to Asia Minor and Europe, and four North American species. It follows, therefore, that this clade diverged from other Pentanthera azaleas at least 40 million years ago, prior to the separation between Western Eurasia and North America (Tiffney, 1985). The demonstration that there is stable and pervasive tetraploidy within clade C does not preclude the sporadic existence of nascent polyloids within primarily diploid species. For example, Rhododendron occidentale was found recently (Jones et al., 2007) to exist both as a diploid in two accessions and as a tetraploid in the cultivar 'Double Dig Twelve'. A preliminary examination of the DNA of 'Double Dig Twelve' showed it to have the same sequences present as in diploid R. occidentale accessions. The occasional tetraploidy of R. occidentale is thus shown to have a more recent origin than that of the species within clade C.
Conclusions
.
The conclusions of our research are:
• Those
Pentanthera
azaleas found consistently as tetraploids make up a monophyletic group that includes
R. luteum
from West Eurasia, as well as four Eastern North American species.
• The Red Hills azalea is a previously undescribed tetraploid species within the tetraploid clade.
• At an early point in the evolutionary divergence of
Pentanthera
azalea species, two lineages separated - one containing the common ancestor of
R. molle
,
R. canadense
and the
tetraploid
clade - the other being the common ancestor of
R. occidentale
and all of the diploid
Pentanthera
species of the Eastern US.
DESCRIPTION OF RHODODENDRON COLEMANII
This species is placed in
Rhododendron
sect.
Penthanthera
G. Don on account of the deciduous leaves, narrow corolla tube, and 5 exserted stamens (Kron, 1993). In the field, the azalea can be distinguished from
R. alabamense
and other coastal azaleas by its late spring (early to mid May) flowering time, its wide range of flower color (white, pink or yellow), its mesic habitats, its ovoid leaves with cuneate bases, its often warty seed capsules with glandular hairs, and its taller stature (3-7 m). Its natural distribution extends from southwestern Alabama to the Chattahoochee Valley in Georgia in areas underlain by strata from the Eocene, Paleocene, and Cretaceous geological periods.

|
|
Figure 4. Typical
Rhododendron colemanii
flowers; flower
and vegetative buds; seed capsule; and plant form, leaves, and habitat--Monroe County, Alabama. |
Rhododendron colemanii R. Miller, sp. nov. TYPE: U.S.A. Alabama: Monroe County, between Highway 41 and Big Flat Creek, 2 May 2007, R. F. Miller 24691 (holotype, TROY). Fig.4. Species nova Rhododendron sectionis Pentanthera distinguibili tetraploideo cum genome mango, multicaulis, deciduo, alabastris elongato, et capsules cum verrucuosis glandulis. Species e habitatine humido, 3-7m elatus, 2C genome amplitude circa 3.16pg DNA, florescentiae May praecox vel medius, flores colores vari, albus, roseus vel luteis, et affinis ad fundamento R. atlanticum , R. austrinum , R. calendulaceum , et R. luteum.
Morphological Description . Many-stemmed shrub or small tree to 7m tall, non-rhizomatous except under stressed conditions; juvenile shoots cinnamon brown, covered with sparse unicellular and sparse to dense (toward the tips) appressed multi-cellular, eglandular hairs. Vegetative bud scales glabrous abaxially; margin densely uni-cellularciliate. Leaves membranaceous and dull green to coriaceous and glossy green, 4-8 x 2-4 cm; blade obovate to occasionally oblong, base cuneate, apex broadly acute, mucronate, entire; adaxial surface and mid-vein sparsely covered with multi-cellular hairs, with unicellular hairs near the petiole; abaxial surface sometimes glabrous, usually covered with scattered uni-cellular and occasional appressed multi-cellular hairs, the abaxial mid-vein with appressed multi-cellular eglandular hairs; margin entire, ciliate with unicellular and appressed multi-cellular eglandular hairs; petioles 0.3-0.7 cm long, densely covered with uni-cellular hairs and appressed multi-cellular eglandular hairs, often with scattered multi-cellular gland-tipped hairs on petioles and the bases of young leaves. Flower buds 1.0-1.5 cm (long) x 0.6-0.8 cm (wide), when immature rather narrow and tapered to a sharp apex. Bud scales awned, brownish-green (drying chestnut brown), lacking dark rims except in winter; adaxial surface villous with unicellular hairs; abaxial surface covered by minute uni-cellular papillae; margin uni-cellular-ciliate, very dense and shaggy toward the tip, non-glandular. Inflorescence appearing after the leaves have expanded; a shortened raceme of 8-10 flowers. Pedicels 0.5-0.7 cm long, densely covered with uni-cellular and multi-cellular glandular hairs. Sepals 0.05-0.1 cm long, margins setose-fimbriate with multi-cellular eglandular hairs; abaxial surface sparsely to densely covered with uni-cellular hairs and multi-cellular eglandular hairs, occasionally with glandular multi-cellular hairs. Corolla white with a yellow blotch on the upper corolla lobe, or uniformly white, pink with yellow blotch, or uniformly pink (rarely yellow); fragrance sweet, musky, and lemony; corolla 2.5-5 cm wide, often reflexed, very variably shaped; corolla tube 1.6-2.5 cm long, 0.25-0.4 cm wide at base; outer surface of corolla covered with uni-cellular and multi-cellular gland-tipped hairs, often in rows on the ridges of the tubes; inner surface of corolla densely covered with uni-cellular hairs. Stamens 4.5-6.7 cm long, with flattened uni-cellular hairs the base of the filament, exserted 2.5-3.5 cm beyond the throat of corolla. Style 5-7 cm long, exserted 3-4 cm beyond the throat, usually curved upward, with dense unicellular hairs on the proximal end, often light brown or reddish toward the distal end; stigma 0.1-0.2 cm wide. Ovary 0.20-0.35 cm long, 0.1-0.2 cm wide at the base, densely covered with uni-cellular and multi-cellular eglandular and occasionally glandular hairs. Fruit a capsule 1.3-3.0 cm long, 0.3-0.5 cm wide at base, cylindrical with prominent sutures and raised warty bumps toward the distal end, sparsely to densely covered with uni-cellular and multi-cellular eglandular hairs, often with glandular-tipped multi-cellular hairs at the proximal end and on the peticels near the sepals, or often scattered along the entire capsule. Seeds pale brown, 1.5-3 mm long; testa expanded, frayed on the ends, and dorsoventrally flattened, surrounding the body, the cells elongate.
Etymology . The epithet honors S. D. Coleman, Sr., of Fort Gaines, Georgia, who collected, propagated, and distributed this azalea to gardens.
|
|
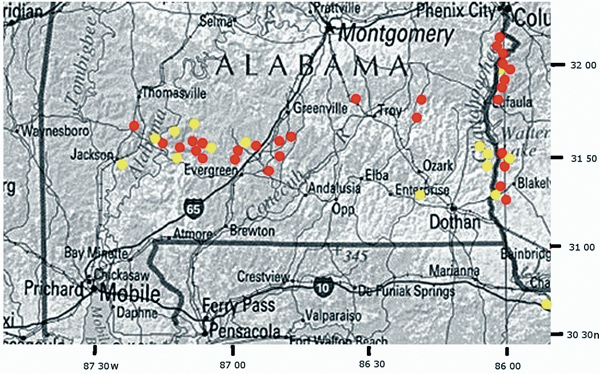
Red =
Rhododendron colemanii
, Yellow =
R. alabamense
Figure 5. Distribution of R. colemanii and R. alabamense in the Coastal Plain of Alabama and Georgia |
Distribution and Ecology
. Southern Alabama and western Georgia in areas underlain by Eocene, Paleocene, and Cretaceous strata - the so-called "Red Hills" (Figure 5) - on well-drained sandy ridges and banks along watercourses or at the base of north-facing bluffs above streams. Associated species:
Rhododendron canescens
(Michaux) Sweet,
Kalmia latifolia
L.,
Stewartia malacodendron
L.,
Hamamelis virginiana
L.,
Fagus grandifolia
Ehrhart,
Pinus glabra
Walter,
R. prunifolium
(Small) Millais [in the east],
Illicium floridanum
Ellis [in the west],
Quercus alba
L.,
Q. laurifolia
Michaux,
Liriodendron tulipifera
L.,
Magnolia macrophylla
Michaux,
M. virginiana
L.,
Cyrilla racemifloria
L.,
Ilex vomitoria
Aiton,
Symplocos tinctoria
(L.) L'Héritier.,
Myrica cerifera
(L.) Small, and
Alnus serrulata
(Aiton) Willdenow. Flowering in early to mid May, occasionally in late April and rarely into June.
Rhododendron colemanii
is superficially similar to
R. alabamense
(Kron, 1993), though they are found in different phylogenetic clades and
R. colemanii
is tetraploid while
R. alabamense
is diploid (Figure 3; Table 1). On the Coastal Plain, both
R. alabamense
and
R. colemanii
bloom after the leaves are exerted; therefore, no distinction can be made on that basis. Their odors are both sweet and somewhat lemony and can be confused. However, compared to
R. alamanense
,
R. colemanii
can also be distinguishing by a later bloom time, its more mesic habitats, longer flower buds, a range of flower colors, and fruit capsules that lack long, appressed cilia but display small, raised warts and glandular hairs (see Table 2 for more detailed comparisons; Figure 4).
Rhododendron canescens
occasionally grows alongside
R. colemanii
, but can easily be distinguished by observing the former plant's rough, elongate leaves; its matted, rusty-colored current-year's growth; its blunt, textured, flower buds; and its late March to early April blooming season.
Rhododendron prunifolium
, often a cohort species within the Chattahoochee drainage, can be identified by its larger bullate leaves; small flower buds; smooth, chocolate-brown current year's growth; and its red to orange, late summer flowers. The range of
R. colemanii
does not overlap with that of
R. eastmanii
Kron and Creel (Kron and Creel, 1999), which blooms at a similar time; the latter azalea occupies mesic sites in South Carolina that are somewhat similar to, though usually steeper and less heavily vegetated than, those occupied by
R. colemanii
.
Rhododendron eastmanii
has a smaller mature size and lacks the range of flower color or multi-stemmed habit of the latter azalea. At least occasionally,
R. eastmanii
displays glands on the edges of the flowerbud scales whereas
R. colemanni
is totally without these glands.
| Table 2. Comparison of distinguishing features between Rhododendron alabamense and R. colemanii . | |
| Rhododendron alabamense | Rhododendron colemanii |
| Colonies in the Red Hills of Alabama and in southwest Georgia bloom from early to mid April | Colonies in the Red Hills bloom from early to mid May. |
| R. alabamense and its hybrids with R. canescens are plants of dry ridges and open, xeric oak woods with scattered Pinus spp., Oxydrendrum arboreum (L.) DC., and Vaccinium arboreum Marsh. | Found primarily on sandy ridges or banks along creeks or on north-facing bluffs in moist woods along with Quercus laurifolia , Pinus glabra Walt., Ilex vomitoria , Cyrilla racemifloria , Magnolia macrophylla , Hamamelis virginiana , Symplocos tinctoria , Myrica cerifera , Kalmia latifolia , and Smilax spp. |
| Though R. alabamense in southern Alabama is a considerably larger plant than the low-growing, stoloniferous plant of Rehder’s type location in northern Alabama, it usually has one or two main stems and an open, flat-topped "umbrella" form when mature. | Usually has 4-8 stems and can become 3-7 m. tall, with a rounded, upright, dense form. |
| Fully developed flower buds are shorter (0.8-1.0 cm) and more rounded in the taper toward the tip. The distal ends of the bud scales, when present, are usually outlined with distinct brown edges. | Flower buds are longer (1.2-1.4 cm) and, when immature, more elongated, without dark rims on the scales except in the winter, and even then rimmed rather lightly. |
| Flowers are typically white, primarily with yellow blotches. The corollas occasionally exhibit eosin pink rims and are thin and almost translucent. | Flowers can be white, pink, or yellow, with or without blotches. The colors often spread across the entire corolla. Flower forms vary from tubular to broadly flared. Flower corollas are thicker and nearly opaque. |
| Seed capsules are slim, cylindrical, with clear sutures, and are covered with unicellular hairs and occasional long, appressed multicellular hairs. Glandular hairs are very rare on the capsules. | Though they have approximately the same shape and size, R. colemanii capsules do not have the long appressed cilia and have surfaces, especially toward the distal end, and are marked by raised warts. Fresh capsules and pedicels, as opposed to material on herbarium sheets, are often covered with mulit-cellular hairs tipped with glands. |
Paratypes . U.S.A. Alabama: Butler County, east of East Chpman at Mashy Creek, 3 May 2007, R. F. Miller 24697 (TROY); U.S.A.: Georgia, Quitman County, Georgetown, River Bluff Park, 4 May 2007, R. F. Miller 24696 (TROY); U.S.A.: Georgia, Quitman County, edge of Soapstone Creek at Highway 39 bridge, R. F. Miller 24696 (TROY).
Literature Cited
Ammal, E.K.J., I.C. Enoch, and M. Bridgwater. 1950. Chromosome numbers in species of Rhododendron.
Rhododendron Year Book
. 5:78-91.
Chamberlain, D.F. and S.J. Rae. 1990. A Revision of
Rhododendron
IV. Subgenus
Tsutsusi
.
Edinb. J. Bot
. 47(2): 89-200.
Greihuber, J., E.M. Temsch, and J.C.M. Loureiro. 2007. Nuclear DNA content measurement, 67-101. In: J. Doležel, J. Greihuber, and J. Suda (eds.).
Flow cytometry with plant cells: Analysis of genes, chromosomes and genomes
. Wiley-VCH, Weinheim.
Goetsch. L.A., A.J. Eckert and B.D. Hall. 2005. The molecular systematics of
Rhododendron
(Ericaceae): a phylogeny based upon RPB2 gene sequences.
Sys. Bot
. 30(3): 616-626.
Jones, J.R., T.G. Ranney, N.P. Lynch, and S.L. Krebs. 2007. Ploidy levels and relative genome sizes of diverse species, hybrids and cultivars of
Rhododendron
.
J. Amer. Rhododendron Soc
. 61(4):220-227.
Kron, K.A. 1993. A revision of
Rhododendron
section
Pentanthera
.
Edinb. J. Bot
. 50: 249-364.
Kron, K.A. and M. Creel. 1999. A new species of deciduous azalea (
Rhododendron
section
Pentanthera
; Ericaceae) from South Carolina.
Novon: A Journal of Botanical Nomenclature
9(3):377-380.
Kurashige, Y., J.I. Etoh, T. Handa, K. Takayanagi, and T. Yukawa. 2001. Sectional relationships in the genus
Rhododendron
(Ericaceae): evidence from
matK
and
trnK
intron sequences.
Plant Systematics and Evolution
228:1-14.
Li, H. Chromosome studies in the azaleas of eastern North America. 1957.
Am. J. Bot
. 44(1):8-14.
Miller, R. and S. Yeatts.
Rhododendron colemanii
: a detective story.
J. Ameri. Rhododendron Soc
. 62:79-85.
Sax, K. 1930. Chromosome stability in the genus
Rhododendron
.
Amer. J. Bot
. 17(4):247-251.
Skinner, H. T. 1955. In search of native azaleas.
Morris Arboretum Bulletin
6:1-2.
Tiffney, B. H. 1985. The Eocene North Atlantic land bridge: its importance in Tertiary and modern phytogeography of the Northern hemisphere.
Jour. Arnold Arboretum
66:243-273.
Towe, L. C. 2004.
American Azaleas
. Timber Press, Portland and Cambridge, 146pp.
Acknowledgements
Recognition and appreciation is given to Alvin Diamond, Buddy Lee, R. O. Smitherman, Bob Stevens, John Thornton, Clarence Towe, and Steve Yeatts for their efforts, insights, and assistance in understanding and describing this species. Thanks to Jeff Jones and Nathan Lynch for completing flow cytometry analyses. We thank Steve Hootman and Rick Peterson for assistance in obtaining samples. Special thanks are given to Dr. Paul Fantz for his review and assistance with the manuscript.
Wenyu Zhou is currently a graduate student, Taylor Gibbons is an undergraduate at the University of Miami, Miami Florida, Loretta Goetsch is a research scientist and Benjamin Hall is a professor of Biology. Thomas G. Ranney is a professor of Horticultural Science and member of the Southeastern Chapter. Ron Miller is currently a retired professor of English literature.